2022 | Self-supervised driven consistency training for annotation efficient histopathology image analysis
https://malmic.ca/wp-content/themes/osmosis/images/empty/thumbnail.jpg 150 150 MaLMIC - Machine Learning in Medical Imaging Consortium MaLMIC - Machine Learning in Medical Imaging Consortium https://malmic.ca/wp-content/themes/osmosis/images/empty/thumbnail.jpg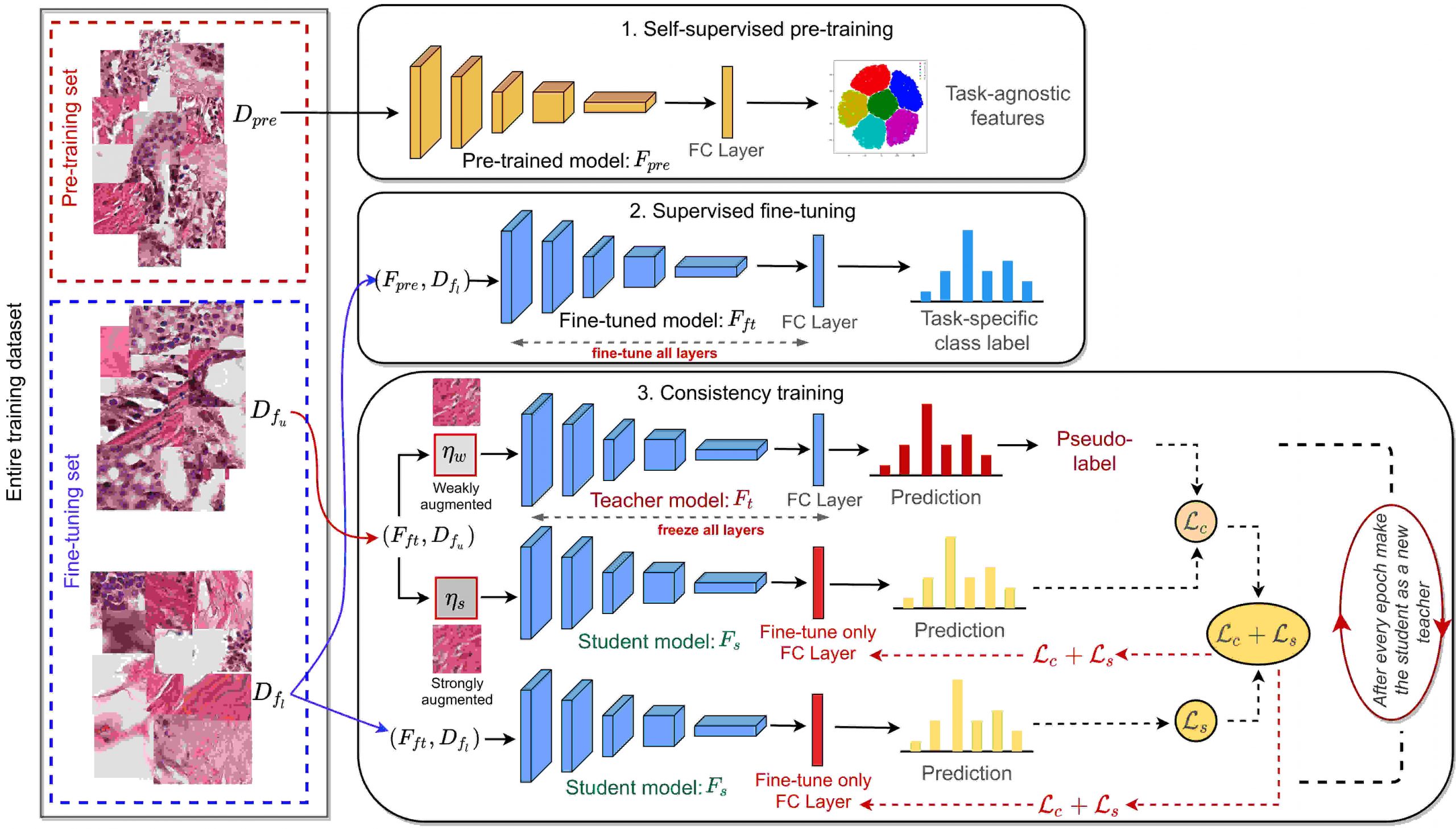




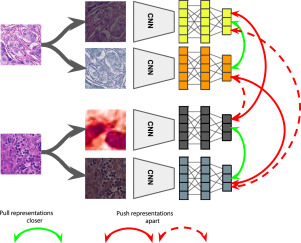
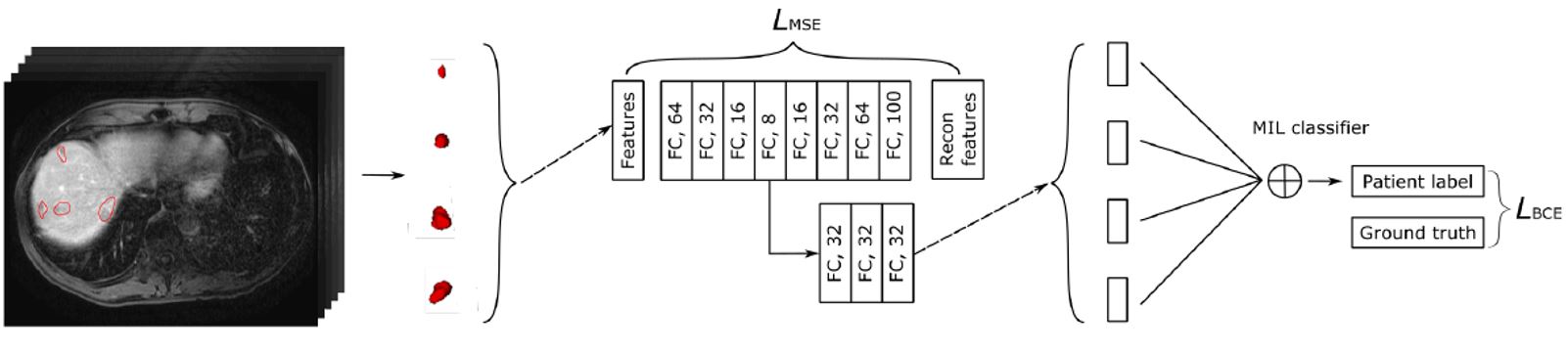
Short Description: In this work, we propose two novel strategies for unsupervised representation learning in histology: (i) a self-supervised pretext task that harnesses the underlying multi-resolution contextual cues in histology whole-slide images to learn a powerful supervisory signal for unsupervised representation learning; (ii) a new teacher-student semi-supervised consistency paradigm that learns to effectively transfer the pretrained representations to downstream tasks based on prediction consistency with the task-specific unlabeled data. We carry out extensive validation experiments on three histopathology benchmark datasets across two classification and one regression based tasks.
Modality: Optical/Microscopy
Organ: Breast
Disease: Cancer
Click Here to Read the Full Publication